Cancer Pathology Data Sharing, published by HL7 International / Orders and Observations. This guide is not an authorized publication; it is the continuous build for version 2.0.0 built by the FHIR (HL7® FHIR® Standard) CI Build. This version is based on the current content of https://github.com/HL7/cancer-reporting/ and changes regularly. See the Directory of published versions
| Page standards status: Informative |
This guide provides an alternative to the HL7 Version 2.5.1 message or HL7 CDA document approach. The following table illustrates the mapping of the well-established NAACCR segments to FHIR resources.
At present, patient and specimen data flow from the clinic and EHR to the pathology lab in a variety of formats, such as HL7 V2 and CDA. The NAACCR Standards for Cancer Registries Volume V, Laboratory Electronic Reporting Pathology Version 5.0, May 2020 (Revised 2020) defines a standard message format for transmitting pathology laboratory information to central cancer registries using the HL7 Version 2.5.1 Message format.
This IG provides an alternative to the HL7 V2 message for promoting better interoperability and scalability. This IG also addresses the collection and exchange of pathology lab data. That is, while discrete data elements are captured in electronic format in most laboratory information systems (LISs), typically, synoptic cancer pathology reports then flow from the LIS to EHRs in non-discrete data formats (e.g., pdf format). This IG helps address these issues.
This IG defines an electronic data exchange standard that maintains the discrete cancer pathology data when exchanging information among pathology laboratories, hospital systems, and registries for continuity of care, structured storage and exchange, standardized cancer reporting, and research. In addition, there is growing industry facilitation of FHIR standards for these kinds of use cases. For instance, starting in December 2022, CMS and ONC will require certified HIT vendors to implement FHIR-based APIs (see ONC’s Cure Act Final Rule). This IG is intended to assist pathology laboratory reporting in moving from V2 to FHIR alignment in accordance with these interoperability policy initiatives.
| V2 Message Elements | NAACCRR Section | FHIR Artifact |
|---|---|---|
| Software Segment (SFT) | 2.15.12 | N/A |
| Patient Identification segment (PID) | 2.6.2 | US Core Patient Profile |
| Patient Visit segment (PV1) | 2.6.2 | US Core Encounter Profile, US Pathology-Related Practitioner, US Core Patient Profile |
| Common Order segment (ORC) | 2.7.1 | US Pathology-Related PractitionerRole |
| Observations Report ID segment (OBR) | 2.7.2 | US Pathology Diagnostic Report, US Pathology-Related Practitioner Role |
| Notes and Comments segment (NTE) | 2.6.4 | US Pathology Diagnostic Report |
| Observation/Result segment (OBX) | 2.7.3 | US Pathology Diagnostic Report results |
| Notes and Comments segment (NTE) | 2.7.4 | US Pathology Diagnostic Report |
| Specimen (SPM) | 2.7.5 | US Pathology Specimen |
| Observation Related to Specimen (OBX) | 2.7.3 | US Pathology Diagnostic Report |
| Continuation Pointer (DSC) |
The IHE Structured Data Capture (SDC) standard, IHE SDC eCP on FHIR, uses a form-driven workflow to capture and encode data and then create FHIR Observations from the captured data. The College of American Pathologists (CAP) has developed and tested the IHE SDC IG for collection of data from electronic Cancer Protocols (eCPs). The goal of IHE SDC eCP on FHIR is to extract data from these cancer checklists and create FHIR Observations for better interoperability between systems.
This Cancer Electronic Pathology Reporting FHIR IG and the IHE SDC eCP on FHIR are aligned and meant to operate together. Pathology data entered into an eCP by a pathologist can be converted into FHIR observations via the IHE SDC on FHIR IG. At that point, it can be transmitted to the Central Cancer Registry (CCR) either directly from the LIS or via an EHR intermediary. Both scenarios are described in this IG.
Figure: Data Flows Supported by this Implementation Guide
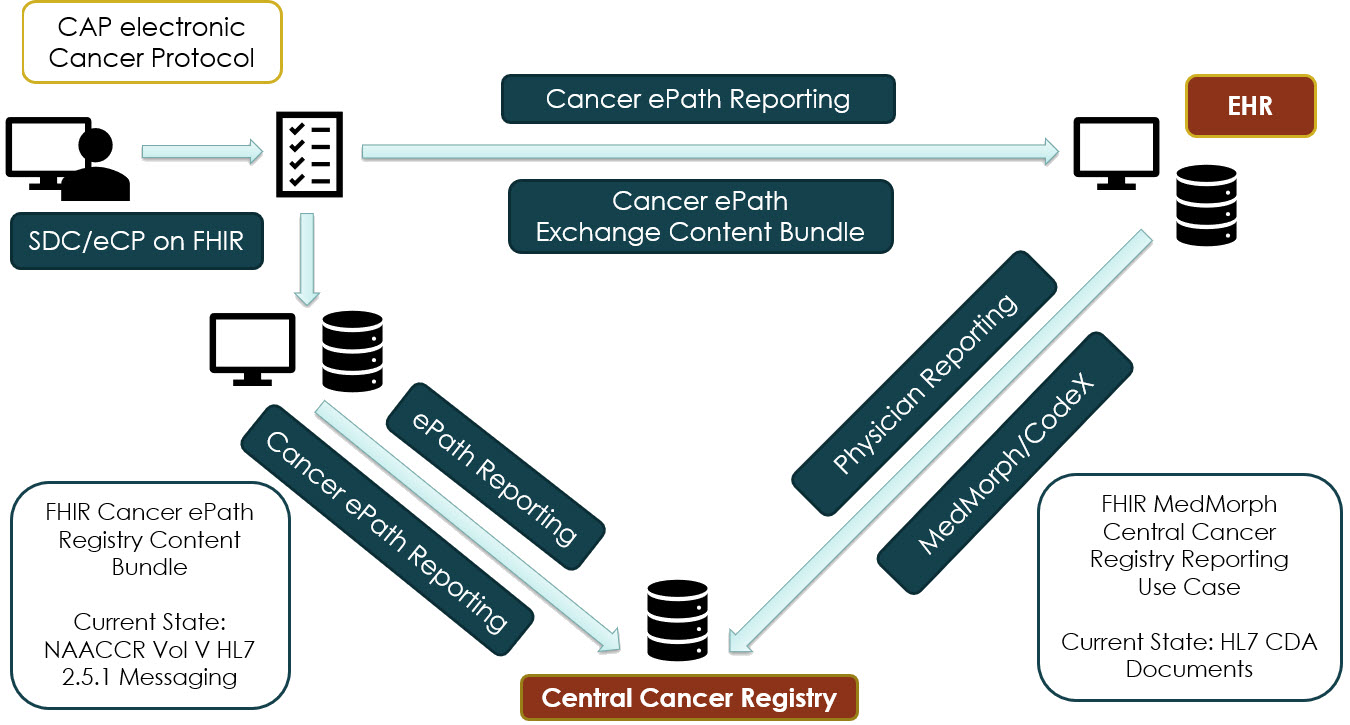 |
This project acknowledges the ongoing discussion about how LOINC vs. SNOMED terminologies serve cancer pathology, but it does not propose to resolve that discussion. Future development of this standard may explore new use cases, such as microbiology laboratory reporting. Future revisions will also include mappings for CAP eCP Protocol CKey Identifier to SNOMED translation.
Several cancer pathology exchange initiatives are underway internationally, including:
Cancer ePathology Reporting Project Contributors
| Role | Name | Affiliation | Contact |
|---|---|---|---|
| Primary Editor | Patrick Joyce | Lantana Consulting Group | patrick.joyce@lantanagroup.com |
| Primary Editor | David deRoode | Lantana Consulting Group | david.deroode@lantanagroup.com |
| Primary Editor | Rick Geimer | Lantana Consulting Group | rick.geimer@lantanagroup.com |
| Co-Editor | Zabrina Gonzaga | Lantana Consulting Group | zabrina.gonzaga@lantanagroup.com |
| Co-Editor | Wendy Wise | Lantana Consulting Group | wendy.wise@lantanagroup.com |
| Co-Editor | Ruby Nash | Lantana Consulting Group | ruby.nash@lantanagroup.com |
| Co-Editor | Diana Wright | Lantana Consulting Group | diana.wright@lantanagroup.com |
| Co-Editor | Max Nakamura | Lantana Consulting Group | max.nakamura@lantanagroup.com |
| Subject Matter Expert | Wendy Blumenthal | Centers for Disease Control and Prevention (CDC) | wfb6@cdc.gov |
| Subject Matter Expert | David Jones | Centers for Disease Control and Prevention (CDC) | pvl1@cdc.gov |
| Subject Matter Expert | Sandy Jones | Centers for Disease Control and Prevention (CDC) | sft1@cdc.gov |
| Subject Matter Expert | Caitlin Kennedy | Centers for Disease Control and Prevention (CDC) | mhy5@cdc.gov |
| Subject Matter Expert | Temitope Alimi | Katmai Government Services contract to CDC | nyj4@cdc.gov |
| Subject Matter Expert | Wendy Scharber | Katmai Government Services/Registry Widgets contract to CDC | wendy@registrywidgets.com |
| Subject Matter Expert | Richard (Rich) Moldwin | College of American Pathologists | rmoldwi@cap.org |
| Subject Matter Expert | Alex Goel | College of American Pathologists | agoel@cap.org |
| Subject Matter Expert | Walter (Scott) Campbell | University of Nebraska Medical Center | wcampbel@unmc.edu |
Current Work Group includes those who participated in the Cancer FHIR IG project: Hans Buitendijk, David Burgess, Lorraine Constable, Robert Hausam, Ralf Herzog, Patrick Loyd, Ulrike (Riki) Merrick, John David Nolen, James Alexander (Alex) Mays, Andrea Pitkus, Craig Newman, James Tcheng, Laura Heermann, and Calvin Beebe.